Biodiversity Reading time 5 min
PAX2GRAPHML: a python library for large-scale regulation network analysis using BioPAX
Published on 08 February 2022

The concept of regulated reactions, which allows connecting regulatory, signaling and metabolic levels, has been used. Biochemical reactions and regulatory interactions are homogeneously described by regulated reactions involving substrates, products, activators and inhibitors as elements.
PAX2GRAPHML is highly flexible and makes it possible to generate graphs of regulated reactions from a single BioPAX source or by combining and filtering BioPAX sources.
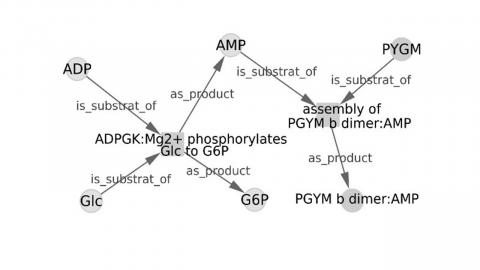
Supported by the graph exchange format .graphml, the largescale graphs produced from one or more data sources can be further analyzed with PAX2GRAPHML or standard Python and R graph libraries.
Availability and implementation: https://pax2graphml.genouest.org.
François Moreews, Hugo Simon, Anne Siegel, Florence Gondret, Emmanuelle Becker, PAX2GRAPHML: a python library for large-scale regulation network analysis using BioPAX, 2021; Bioinformatics, 37 (24): 4889-4891. https://doi.org/10.1093/bioinformatics/btab441
